| Set | A set includes a particular list of compounds; multiple sets can be present at once and are displayed on individual tabbed panes in the GUI. |
| Project | A project comprises all data and compound sets when running an instance of cApp. In the graphical user interface (GUI), each compound set is displayed as a table on an individual tab. Automatically generated HTML, PDF and ASCII presentations of compound sets are identified by their set number. |
| Task | A cheminformatics task that is to be performed by the program. Different tasks result in different visual representation. |
| Appraise | Physico-chemical properties and structural features will be calculated and analysed as to compliance with various likeness criteria and existence of PAINs components. User-provided data and annotation can be included and interactive convenience features are available. |
| Similarity search | Libraries of small-molecule compounds can be queried using individual or multiple compounds for structural similarity with a maximum common subgraph approach. The PubChem Compound database can also be queried for similarity. |
| Compound clustering | Using Tanimoto similarity, the compounds within one set or grouped in to a user-specified number of clusters. |
| Splitting of libraries | Multi-compound files can be split into subsets with a user-specified number of entries each. Currently only possible for libraries in SDF format. |
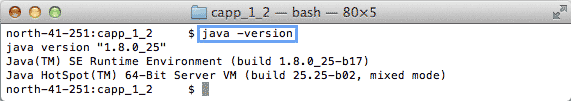
| 1. | The Java Runtime Environment (JRE) has to be at least Java 1.7. Use the following command to check your JRE version: java -version |
 | 2. | Navigate to the directory in which the cApp jar file resides using the following command: cd {folder} |
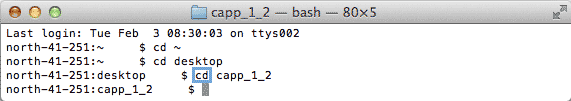 |
| 3. | To ensure that the correct file is present in the directory, enter ls |
 |
| 4. | Enter the following command to start the program with default settings (i.e. with the GUI): java -jar capp_v1.5.1_java1.7.jar |
 |
| 5. | On Linux, a shell script can be used to access the program conveniently. | #!/bin/csh -f setenv JAVA {path-to-java-bin-directory}/java setenv CLASSPATH {path-to-capp-executable}/capp_v1.5.1_java1.7.jar $JAVA capp $* |
| Switch | Function |
| -appraise | Runs the appraise task (default) |
| -ascii | Write results in ASCII format (directory: capp_results_ascii) |
| -autoselect | Auto-select largest entity when reading SDF or SMILES |
| -cluster {N} | Group compounds into {N} clusters |
| -debug | Debug option |
| -drug | Evaluate for drug likeness (Lipinski's Rule of 5) |
| -duplicates inchi | Check for duplicate entries based on InChI Keys |
| -duplicates tanimoto | Check for duplicate entries based on Tanimoto similarity |
| -fragment | Evaluate for fragment likeness (Rule of 3) |
| -gui | Start the program with the graphical user interface |
| -h | Print help |
| -html | Generate results in HTML format (directory: capp_results_html) |
| -i {input file} | Input file with compounds to process |
| -inchi | Input file contains InChI code |
| -lead | Evaluate for lead likeness |
| -load | Load a previously saved cApp project |
| -maxsets {N} | Maximum number of compound sets in the project (default: 10) |
| Write results into PDF files (directory: capp_results_pdf) | |
| -pdf landscape | PDF paper orientation is Landscape (default) |
| -pdf portrait | PDF paper orientation is Portrait |
| -pdf bondlength | Structure images are drawn with same bond length (default) |
| -pdf fixed | Structure images have the same fixed size |
| -png | Write PNG images of compounds (directory: capp_images) |
| -pubchem | Search for entry in PubChem when conducting the appraise task |
| -sdf | Input file is an SDF file |
| -smi | Input file contains SMILES code |
| -smsd {library file} | Similarity search of {input file} against {library file} in SDF format |
| -split {N} | Split an SDF library into subsets of {N} entries each |
| -svg | Write SVG images of compounds (directory: capp_images) |
| -3d | Attempt to generate 3D coordinates |