| 1. | Click File in the menu bar, then click Load project. | 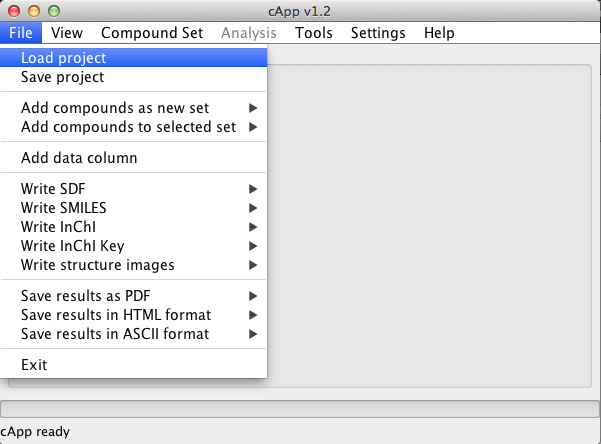 |
| 2. | Binary cApp files should, but don't have to carry a '.cpp' extension. This format re-stores all data from a previous user session (i.e. an entire project). | 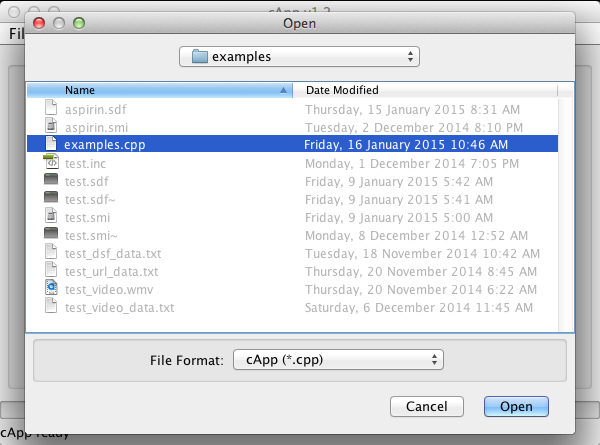 |
| 3. | To save the current project, use File - Save project. |  |
| 1. | Adding compounds as a new set is generally the first step undertaken to begin processing compounds. | 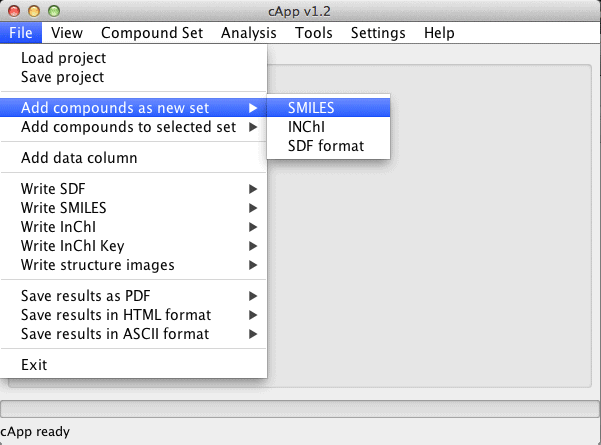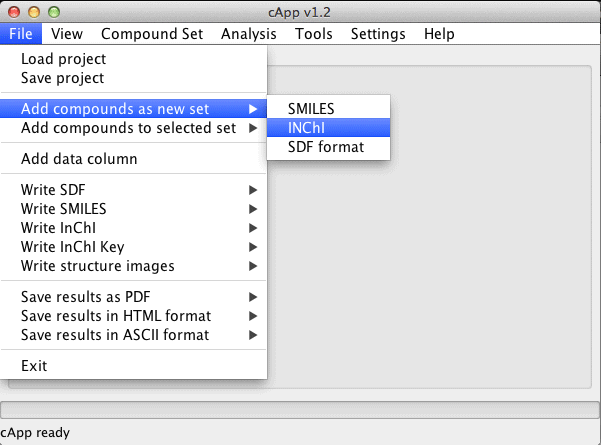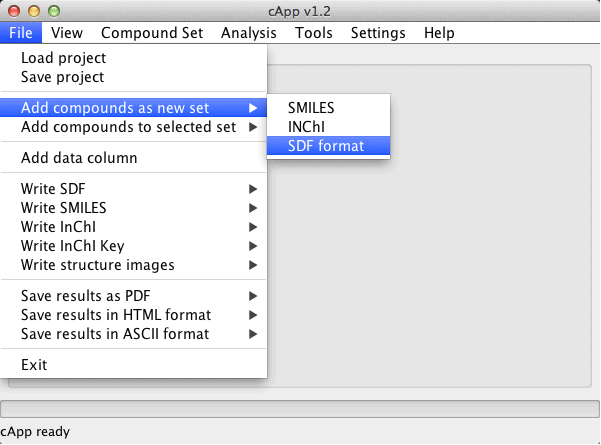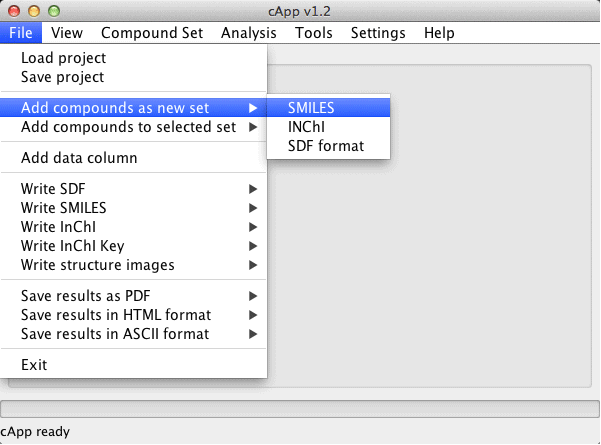 |
| 2. | From the menu bar, select File - Add compounds as new set. In the example here, SDF format has been selected. | 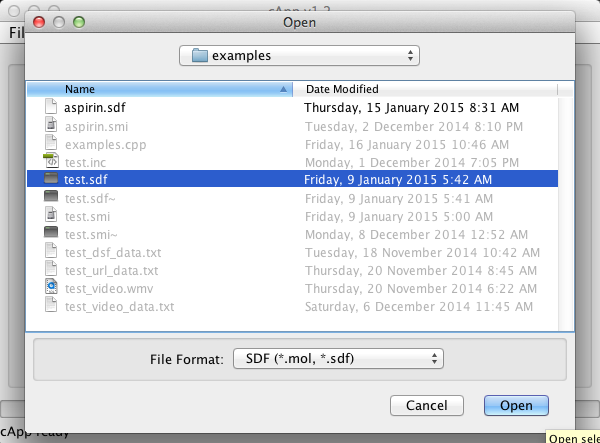 |
| 3. | The file will be loaded and the appraise task will automatically be conducted. |  |
| 4. | The loaded compounds will be added to the project as a new set and displayed as table in a tabbed panel. Sets will be numbered consecutively. The default name displayed is the file name; however, this can be changed using Compound Set - Compound set description from the menu bar. | 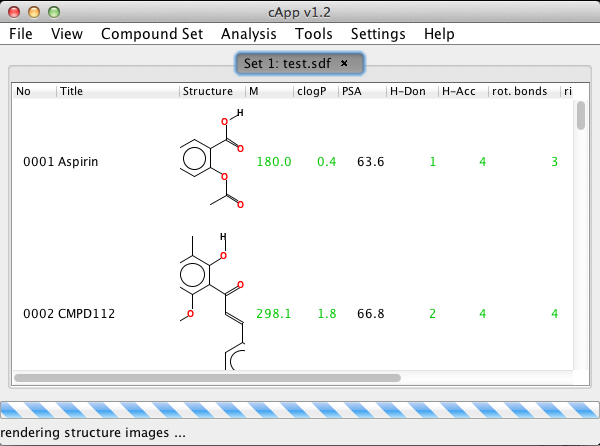 |
| Compounds can also be added to an existing set. This set needs to be selected in the GUI (click on the tab at the top of the panel with the desired set). Then choose File - Add compounds to selected set from the menu bar. | 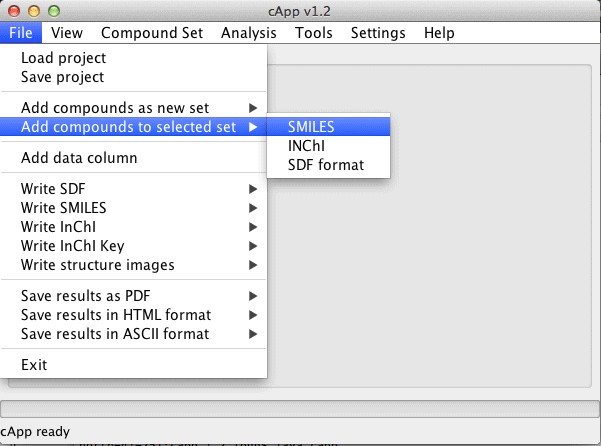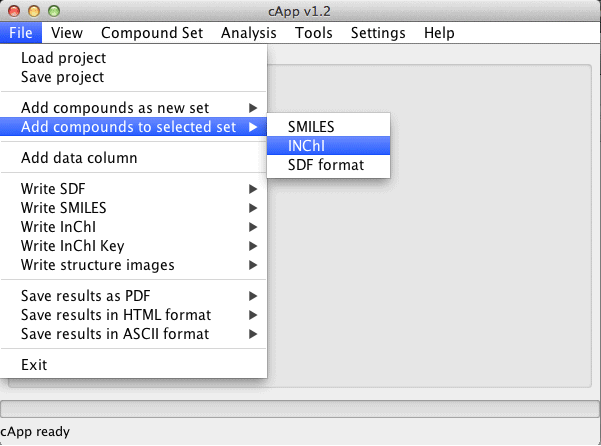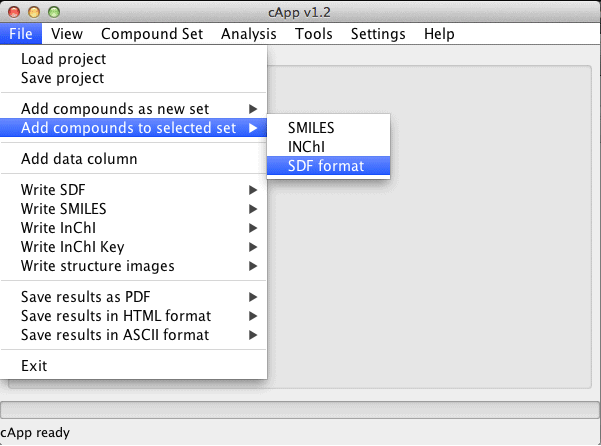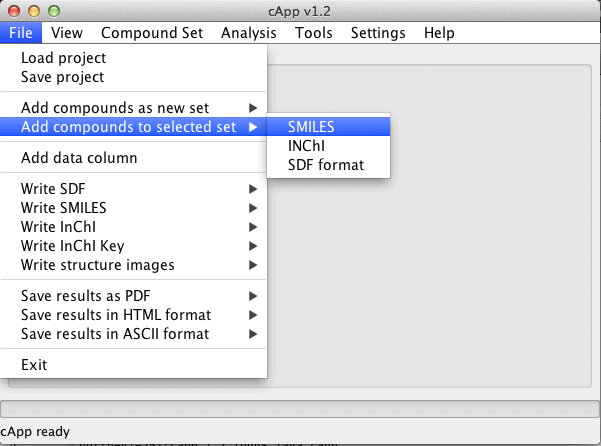 |
| 1. | Select the compound of interest by a left-click in that row. | 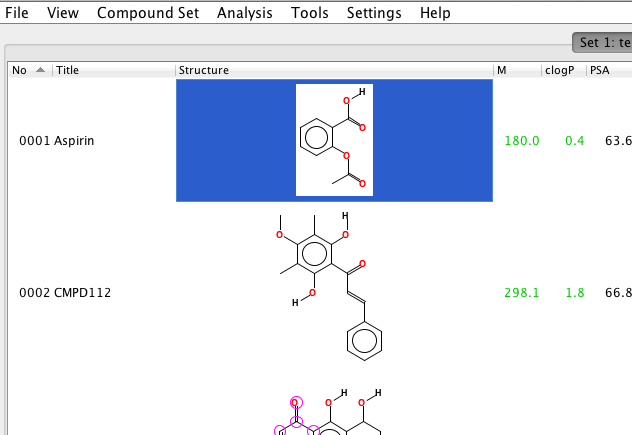 |
| 2. | A right-click opens the pop-up menu; select Show meta data. | 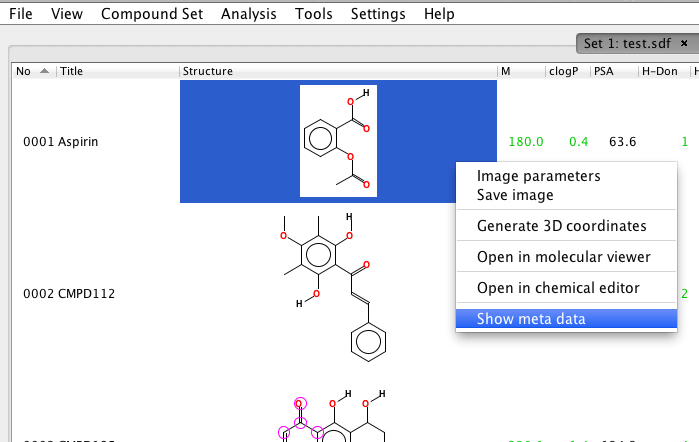 |
| 3. | The meta data for the selected compound are displayed in a new window (and can be copied via the clipboard). | 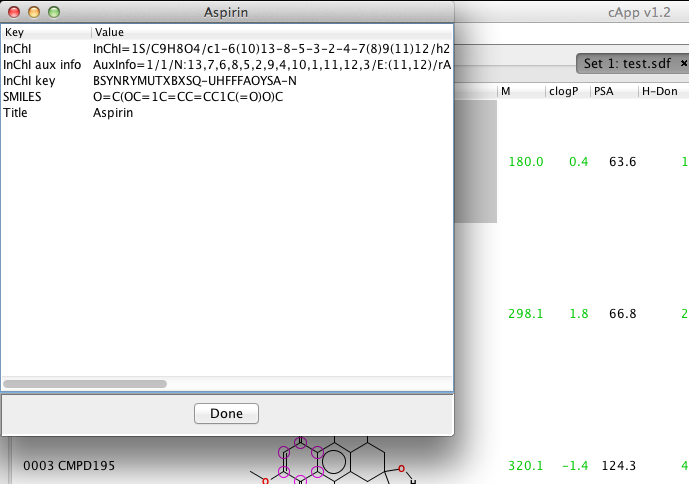 |
| 4. | Optional (this is not cApp-specific): meta data may be edited in the SDF file using an ASCII text editor. | 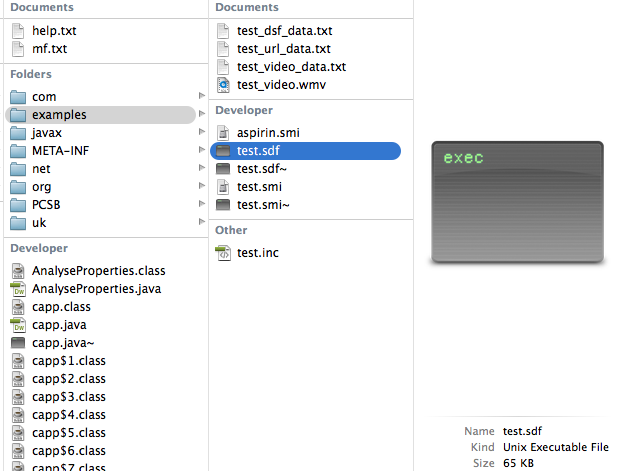 |
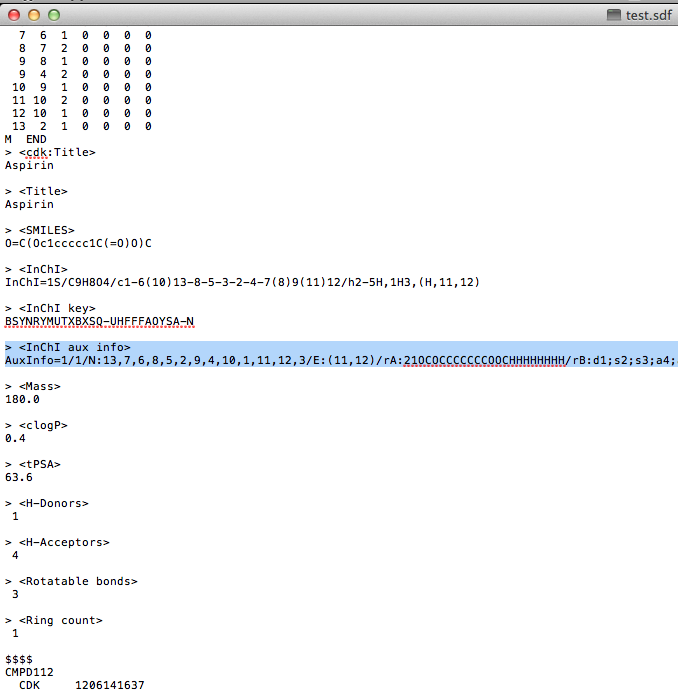
| 5. | Optional (this is not cApp-specific): open the SDF file in a text editor (e.g. textEdit, Notepad etc.) for editing. The key-value syntax for adding properties is as follows: > <Key>
Value |
 |
| 1. | In order to conduct a search for all compounds of a set, select Tools - Identical entries in PubChem - For active set from the menu bar. | 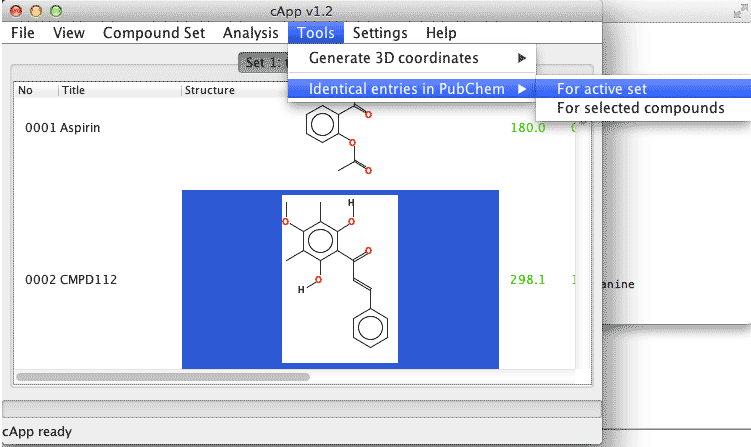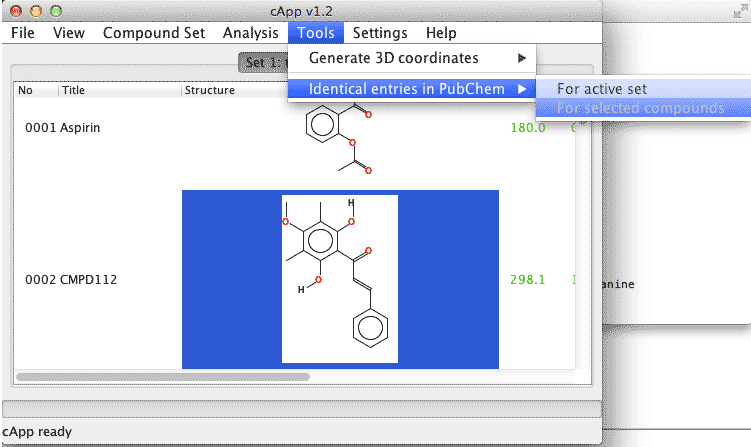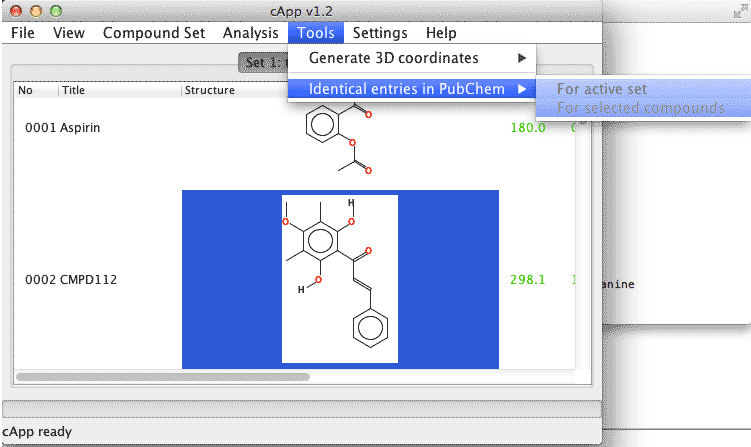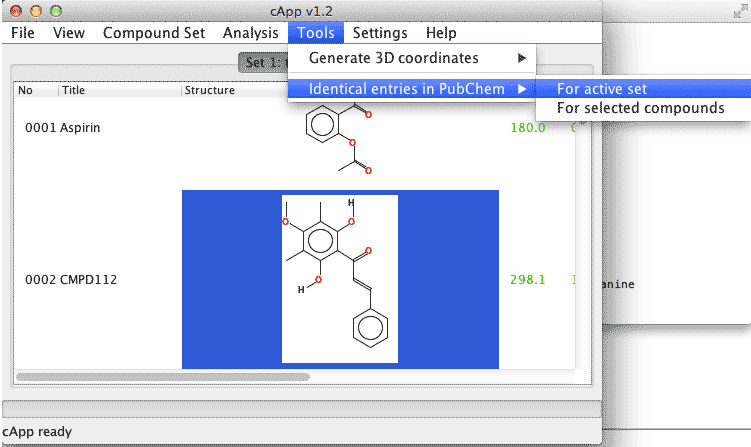 |
| 1. | Select the query compound by a left-click (not in the structure column, though). Then use a right-click to get the pop-up menu and select Find PubChem entry. | 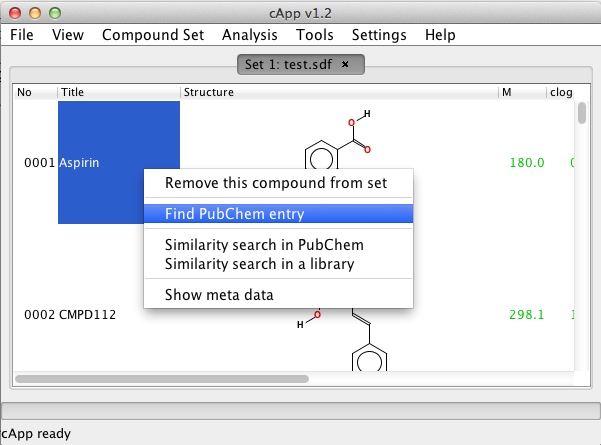 |
| 2. | In the column PubChem CID, a URL will appear linking to the first found entry in the PubChem Compound Database. The entry can be accessed by a left-click, which will display the content in the user's web browser. If no entry is found, the label 'none' will appear in the PubChem CID column, to indicate that this compound has been processed. | 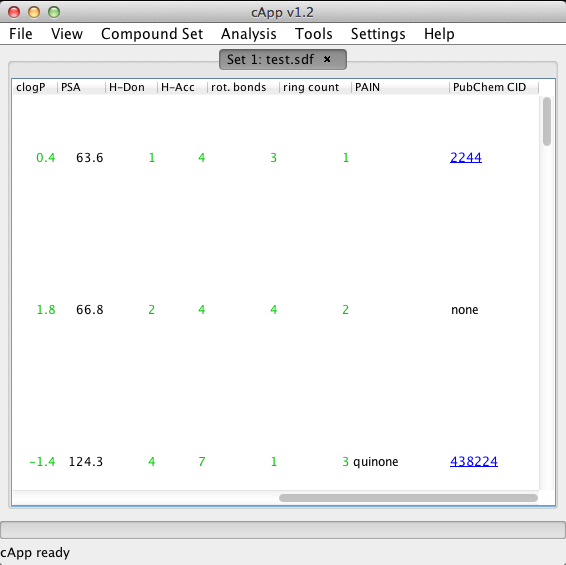 |
| For selected compound | Select the compound by a left-click and obtain the pop-up menu by a right-click. Then choose Generate 3D coordinates. | 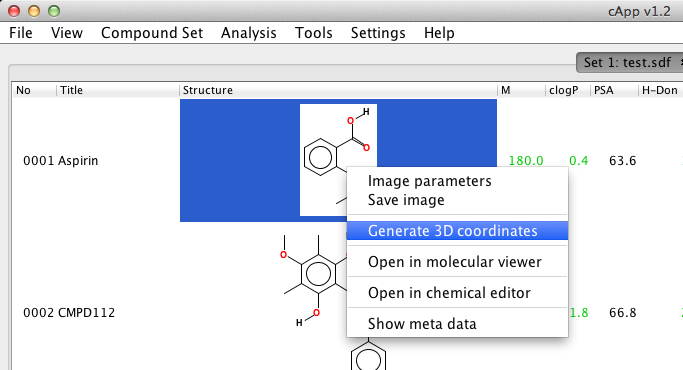 |
| For all compounds in a set | From the menu bar, choose Tools - Generate 3D coordinates to generate 3D coordinates for all compounds in the active set. The calculation will be done in the background as indicated by the progress bar. It is possible to abort the calculation by clicking the button Stop 3D coordinates generation above the tabbed panel. | 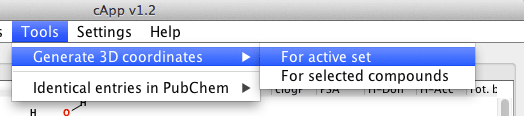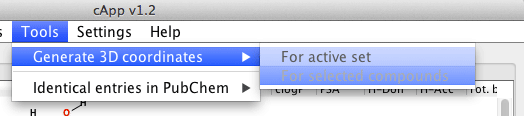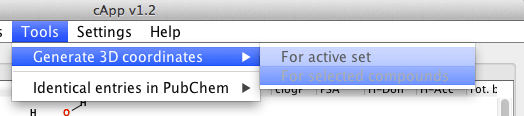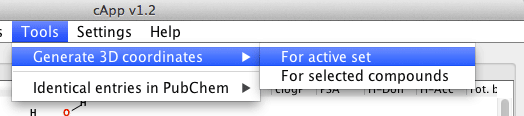 |
| For selected compound | The three-dimensional coordinates of the selected compound can be updated by loading information from a user-provided SDF file. This feature does not affect any other properties of the compound and just updates the atomic coordinates stored for this compound in the SDF component of the entry. This feature is available from the pop-up menu at Update 3D coordinates from external SDF file. When loading an external SDF file, cApp will perform a check to validate that the imported molecule matches the compound entry. For this check, the unique SMILES codes of compound entry and new molecule are compared. |
| The molecular viewer window is available from the pop-up menu (right-click) for the selected compound. The display of hydorgen atoms in the viewer depends on the settings for the selected compound (available in the feature Image parameters from the pop-up menu). | 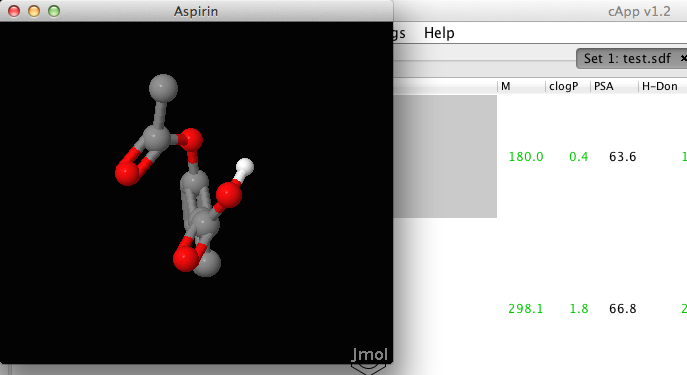 |